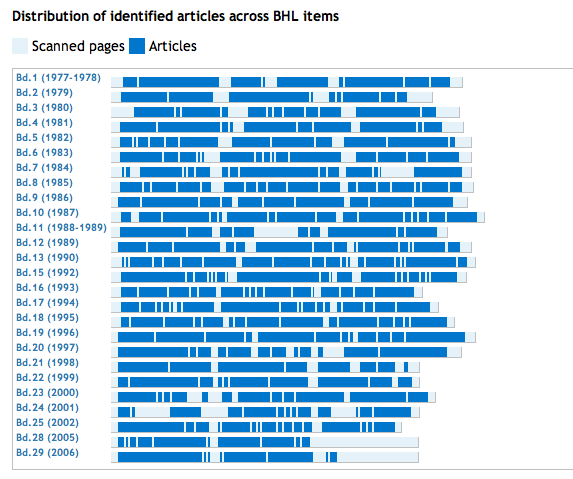
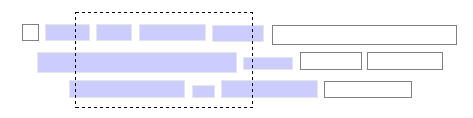
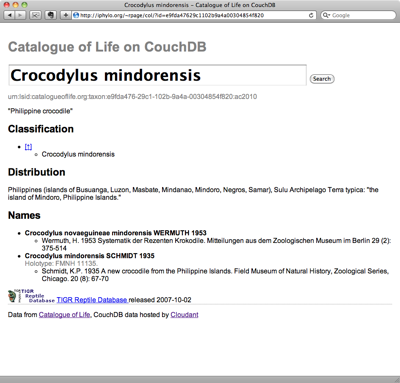
Those who seek answers to big, broad questions about biology, especially questions emphasizing the organism (taxonomy, evolution and ecology), will soon benefit from an emerging names-based infrastructure. It will draw on the almost universal association of organism names with biological information to index and interconnect information distributed across the Internet. The result will be a virtual data commons, expanding as further data are shared, allowing biology to become more of a ‘big science’. Informatics devices will exploit this ‘big new biology’, revitalizing comparative biology with a broad perspective to reveal previously inaccessible trends and discontinuities, so helping us to reveal unfamiliar biological truths. Here, we review the first components of this freely available, participatory and semantic Global Names Architecture.Do we need names?
Reading this (full disclosure, I was a reviewer) I can't wondering whether the assumption that names are key really needs to be challenged. Roger Hyam has argued that we should be calling time on biological nomenclature, and I wonder whether for a generation of biologists brought up on DNA barcodes and GPS, taxonomy and names will seem horribly quaint. For a start, sequences and GPS coordinates are computable, we can stick them in computers and do useful things with them. DNA barcodes can be used to infer identity, evolutionary relationships, and dates of divergence. Taken in aggregate we can infer ecological relationships (such as diet, e.g., doi:10.1371/journal.pone.0000831), biogeographic history, gene flow, etc. While barcodes can tells us something about an organism, names don't. Even if we have the taxonomic description we can't do much with it — extracting information from taxonomic descriptions is hard.
Furthermore, formal taxonomic names don't seem terribly necessary in order to do a lot of science. Patterson et al. note that taxa may have "surrogate" names":
Surrogates include provisional names and specimen, culture or strain numbers which refer to a taxon. 'SAR-11' ('SAR' refers to the Sargasso Sea) was a surrogate name given in 1990 to an important member of the marine plankton. Only a decade later did it become known as Pelagibacter ubique.
The name Pelagibacter ubique was published in 2002 (doi:10.1038/nature00917), although as a Candidatus name (doi:10.1099/00207713-45-1-186), not a name conforming to the International Code of Nomenclature of Bacteria. I doubt the lack of a name that follows this code is hindering the study of this organism, and researchers seem happy to continue to use 'SAR11'.
So, I think that as we go forward we are going to find nomenclature struggling to establish its relevance in the age of digital biology.
If we do need them, how do we manage them?
If we grant Patterson et al. their premise that names matter (and for a lot of the legacy literature they will), then how do we manage them? In many ways the "Names are key to the big new biology" paper is really a pitch for the Global Names Architecture or GNA (and it's components GNI, GNITE, and GNUB). So, we're off into alphabet soup again (sigh). The more I think about this the more I want something very simple.
Names
All I want here is a database of name strings and tools to find them in documents. In other words, uBio.
Documents
Broadly defined to include articles, books, DNA sequences, specimens, etc. I want an database of [name,document] pairs (BHL has a huge one), and a database of documents.
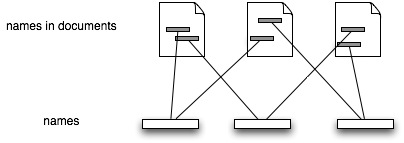
Realistically, given the number and type of documents there will be several "document" databases, such as GenBank and GBIF. For citations Mendeley looks very promising. If we had every taxonomic publication in Mendeley, tagged with scientific names, then we'd have the bibliography of life. Taxonomic nomenclators would be essentially out of business, given that their function is to store the first publication of a name. Given a complete bibliography we just create a timeline of usage for a name and note the earliest [name,document] pair:
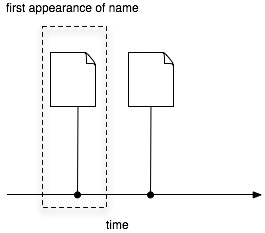 Taxonomy
TaxonomyThere are a few wrinkles to deal with. Firstly, names may have synonyms, lexical variants, etc. (the Patterson et al. paper has a nice example of this). Leaving aside lexical variants, what we want is a "view" of the [name,document] pairs that says this subset refer to the same thing (the "taxon concept").
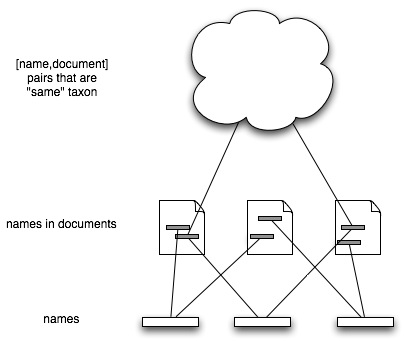
We can obsess with details in individual cases, but at web-scale there are only two ones that spring to mind. The first is the Catalogue of Life, the second is NCBI. The Catalogue of Life lists sets of names and reference that it regards as being the same thing, although it does unspeakable things to many of the references. In the case of NCBI the "concepts" would be the sets of DNA sequences and associated publications linked to the same taxonomy id. Whatever you think of the NCBI taxonomy, it is at least computable, in the sense that you could take a taxon and generate a list of publications 'about" that taxon.
So, we have names, [name,document] pairs, and sets of [name,document] pairs. Simples.